Analysis of BacMam Transduced Hela Cells with High-Throughput Live-Cell Imaging
Key Findings:
BacMam transduction of mammalian cells was optimized using the CytoSMART Omni FL. The transduction efficiency was affected by the virus concentration, cell density, and incubation time. BacMam transduction reduced the proliferation of HeLa cells in a dose-dependent manner. The most optimal conditions of 40 PPC and 20,000 cells/well resulted in high GFP expression and low toxicity.
Abstract
Transduction is a routine method to introduce foreign genetic elements into eukaryotic cells. The BacMam gene delivery system has been widely used to deliver and (over)express genes in mammalian cells. While this technology enables fast and convenient analysis to be performed in a variety of cells, an ideal transduction protocol needs to be established to achieve optimal transgene transduction efficiency. In this application note, the CytoSMART Omni FL live-cell imager was used to assess transduction efficiency and optimize gene expression in HeLa cells. The cells were transduced with varying BacMam Nucleus-GFP concentrations and at different cell densities. Furthermore, the toxicity of the BacMam particles was evaluated through the confluence level measurements. Subsequently, we determined the optimal transduction conditions resulting in high transduction efficiency and minimal cell toxicity.
Introduction
Transduction, like transfection, is a process involving the introduction of exogenous nucleic acids into eukaryotic cells and is used in research to study and modulate gene expression. Unlike transfection, transduction introduces genetic material using a viral vector. This powerful tool allows researchers to better understand the functional significance of genes and gene products by enhancing or inhibiting specific gene expression in cells. By doing so, transduction-based assays can shed more light on cell physiology and the complex molecular mechanisms involved in diseases [1].
Transfection, as well as transduction, can be classified into two types: the foreign genetic material can be stably integrated into the genome of host cells or transiently maintained in the nucleus. A stable transfection leads to permanent genetic changes, whereas a transient transfection results in temporary gene expression effects. As such, a transient transfection and transduction is an ideal, rapid method to investigate the short-term impact of altered gene or protein expression [2].
Numerous approaches have been developed to facilitate the transfer of nucleic acids into cells, including physical, chemical, and viral strategies [2]. Among the viral methods, the BacMam virus expression system has been commonly used. This technology uses a baculovirus – an insect virus modified with a recognizable mammalian promoter – to efficiently deliver and (over)express genes in mammalian cells. The method enables safe, rapid, and facile transduction experiments to be performed in a variety of mammalian cells, including cell lines [3], primary cells [4], and stem cells [5]. Therefore, the BacMam system has been exploited for a growing number of in vitro applications, including the modification and insertion of genes [6], vaccine development [7], and fluorescent labeling of cellular organelles involving the transfer of green fluorescent protein (GFP) fusion constructs [8].
Like any other transduction method, the BacMam technology does not transduce each cell type with equal efficiency, thus the transduction conditions need to be optimized per cell type and BacMam construct. The transduction efficiency can be easily assessed in realtime with fluorescence microscopy when the genetic product is fused to a fluorescent protein. A commercially available format of BacMam (Invitrogen CellLightsTM) contains fusion constructs of fluorescent proteins with signal peptides that provide accurate targeting to specific cell structures (e.g. nucleus). These constructs are taken up by the cell via endocytosis and the released DNA is translocated to the nucleus for transcription (Fig. 1). Following gene expression, the fluorescent proteins can be detected within 24 hours. The CytoSMART Omni FL can be used to monitor the expression of these proteins, while examining multiple transduction conditions simultaneously.
In this application note, we demonstrate how the CytoSMART Omni FL can be used to optimize BacMam transduction conditions. HeLa cells were transduced with BacMam Nucleus-GFP constructs, expressing GFP fused to the SV40 nuclear localization sequence. We evaluated the effects of virus concentration and cell density on the GFP expression. Additionally, we investigated possible toxic effects of the BacMam particles on cellular growth by examining the confluence levels over time using the integrated confluence software.
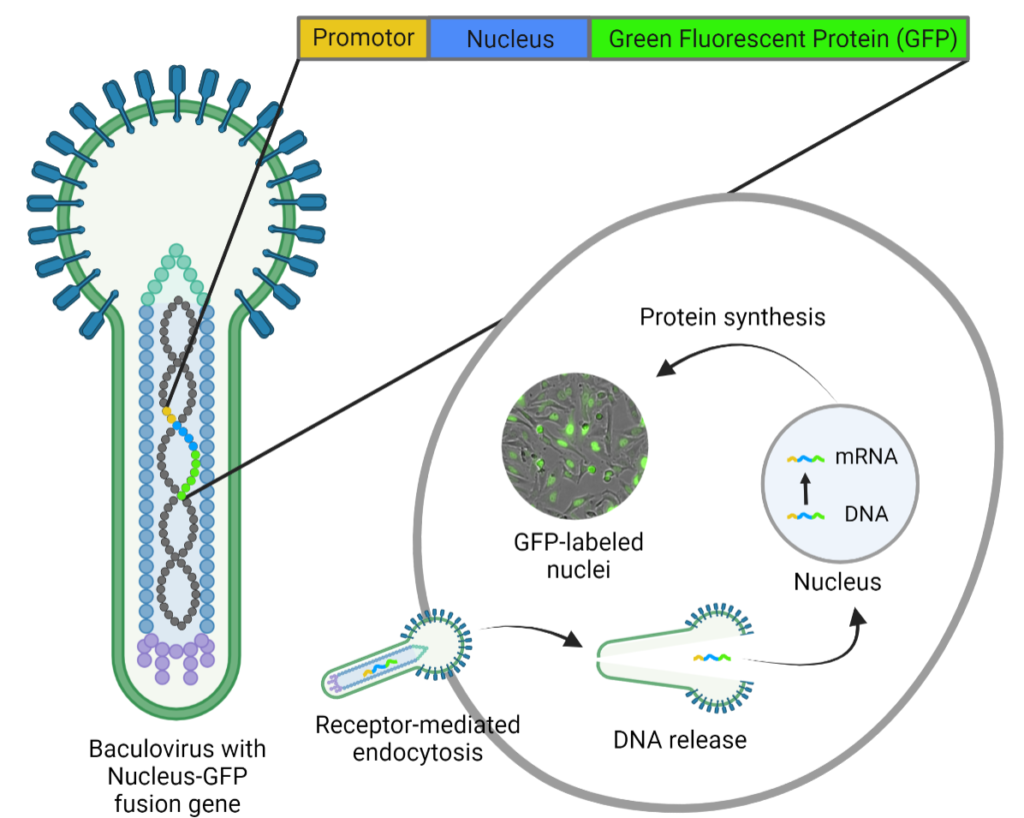
Figure 1: BacMam-mediated gene delivery. The baculovirus carries DNA of the mammalian promoter and target protein coupled to a fluorescent protein (e.g. Green Fluorescent Protein). BacMam particles are taken up by the cell via endocytosis, after which the DNA is translocated to the nucleus for gene expression. Image modified from [9].
Materials and Methods
Cell Culture and Seeding
HeLa cells were cultured to sub-confluency in Advanced DMEM (Gibco) supplemented with 10% fetal bovine serum (Gibco) and 1% Penicillin/streptomycin (Gibco). Cells were plated in a 96-well, clear-bottom black microplate (Thermo, cat. 237105) at densities of 10,000, 20,000, or 30,000 cells per well (100 µL). The plate was then incubated at 37°C 5% CO2 for 4 h.
BacMam Virus Incubation
CellLight Nucleus-GFP, BacMam 2.0 (Thermo, cat. C10602) was added to the cells in five different concentrations: 20, 40, 60, 80, and 100 particles per cell (PPC) and each concentration treatment was assessed in triplicate (Fig. 2A). The working volume was kept consistent in all wells by addition of pre-warmed PBS (Gibco).
Fluorescence Live-Cell Imaging
The 96-well plate was placed on the CytoSMART Omni FL inside an incubator (37°C, 5% CO2). For each condition, brightfield (exposure: 10 ms) and green fluorescence images (exposure: 300 ms, gain: 25, intensity: 75%) were acquired every hour for 72 h.
Evaluation of Transduction Efficiency and Confluency
The efficiency was determined for each condition using ImageJ software (version 1.53). The brightfield images were used to determine the total number of cells and the green fluorescence images were used to count the number of GFP-positive nuclei. The transduction efficiency (%) was then calculated according to (the number of GFP-positive nuclei / total number of cells) x 100. The brightfield confluence levels were tracked over time using the integrated confluence software. Confluence levels were normalized to the cell density at the start of transduction.
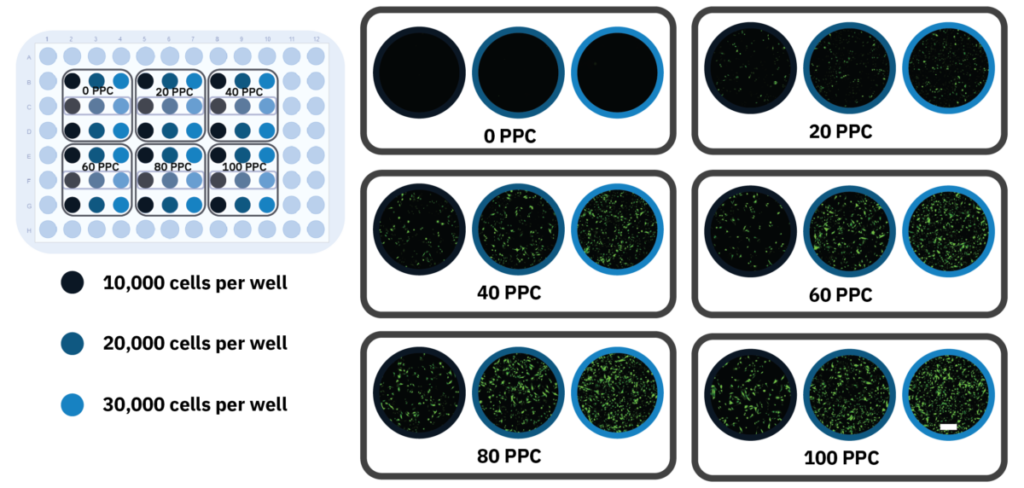
Figure 2: High-throughput optimization of transduction conditions. A) Schematic overview of a 96-well plate indicating the different plating densities and virus concentration treatments. B) Fluorescence images of the different conditions taken after 24 h incubation, showing Nucleus-GFP expression. The outlined wells showcase individual replicates. The scale bar indicates 200 μm.
Results
Nuclear GFP Expression could Be Observed with the Omni FL
Nuclear staining, as compared to actin staining, provides a more efficient approach of evaluating transduction efficiency. After an incubation period of 24 h, images of HeLa cells infected with various BacMam concentrations at different seeding densities confirmed nuclear expression of GFP in nearly all transduced samples (Fig. 2B). A trend can be observed that the GFP expression increased with higher viral load (PPC). Interestingly, GFP expression was concentrated in the nucleus for the lowest virus concentration, whereas the GFP signal was more intense and also located outside the nucleus for the higher concentrations, indicating over-expression.
Virus Concentration Affected the Percentage of GFP Expressing
To evaluate the transduction efficiency, we first determined the effect of the virus concentration on the percentage of GFP expressing cells. The quantitative measurements confirmed the concentration-dependent expression (Fig. 3A). The highest transduction rate was achieved with a concentration of 100 PPC, in which 94.5% of the cells were transduced after 31 h. Notably, the percentage of GFP-expressing cells decreased after this period for 80 and 100 PPC.
Virus Concentration-Dependent Influence on the Cell Proliferation
Parallel analysis of cell proliferation was performed to determine whether the transduction affected the proliferation (Fig. 3B). It can be seen that the cells of all transduced samples proliferated equally up to 18 hours, after which different growth kinetics can be observed.
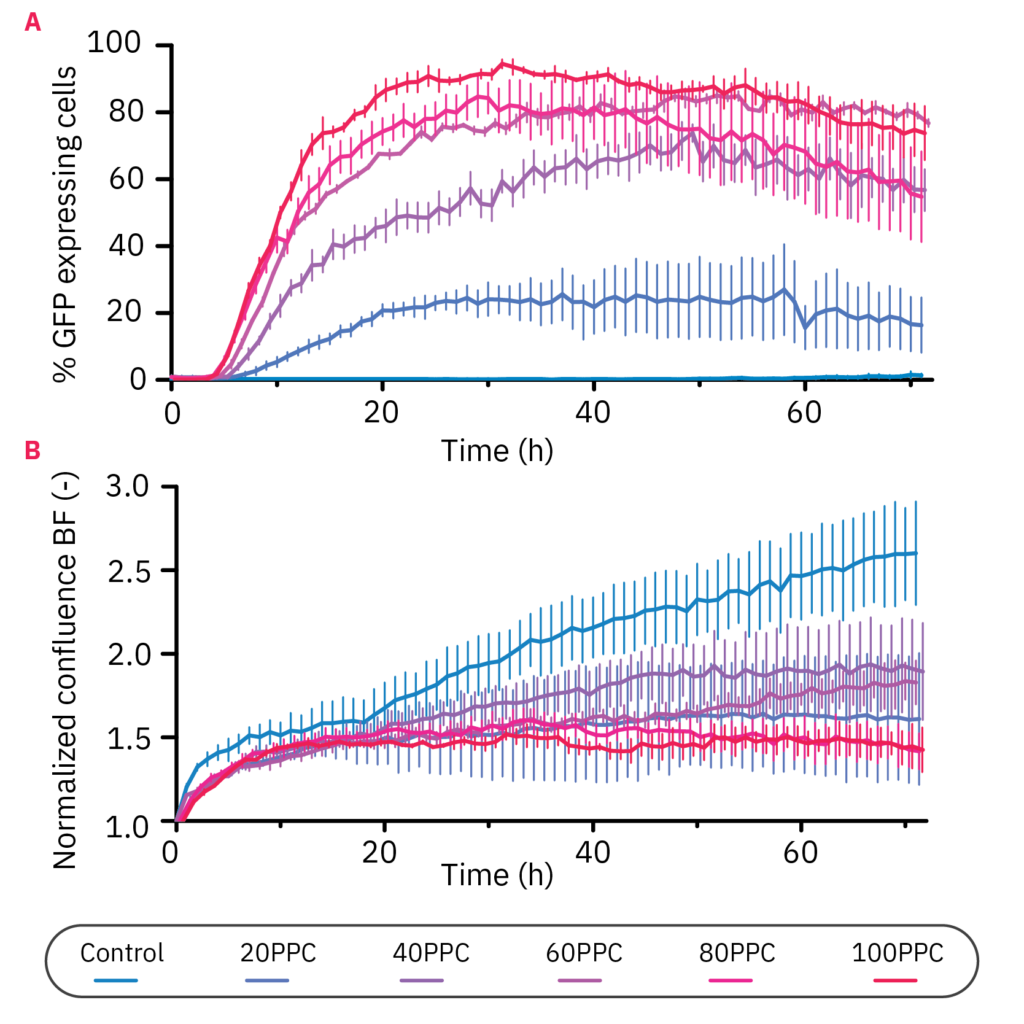
Figure 3: Virus concentration affected GFP expression and HeLa proliferation levels. A) The percentage of GFP-expressing cells over time in 10,000 HeLa cells transduced with different virus particle per cell (PPC) concentrations. The cells were monitored for 72 h with the CytoSMART Omni FL. B) Brightfield confluence levels of 10,000 cells transduced with different virus particle concentrations. Data is displayed as the mean value of 3 replicates each with SEM.
The control group continued to proliferate exponentially, whereas the cells at a concentration of 20, 40, and 60 PPC proliferated at a slower rate, each with slightly different kinetics. Interestingly, the samples with 80 and 100 PPC showed a reduction in confluency over time.
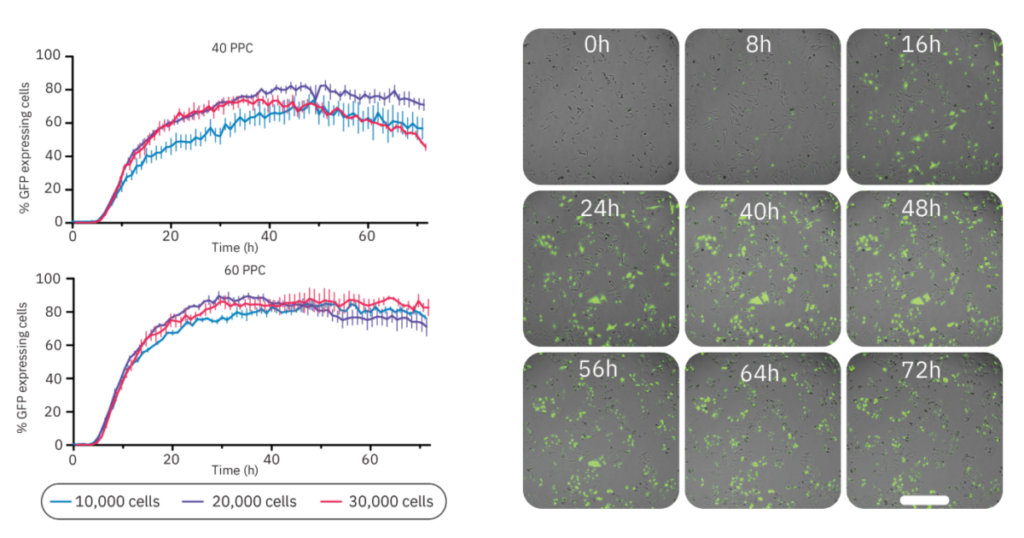
Figure 4: GFP expression was dependent on cell density and virus concentration. (Left) The percentage of GFP-expressing cells infected at different cell densities (10,000, 20,000, or 30,000 cells/well) and with different virus concentrations (40 PPC or 60 PPC). Data is displayed as the mean value of 3 replicates each with SEM. (Right) Time-lapse images of BacMam-mediated transduction of HeLa cells. HeLa cells were transduced at 20,000 cells/well with a concentration of 40 PPC. Upon transduction, Nucleus-GFP fusion protein is expressed in cell nuclei. The scale bar indicates 200 µm.
Cell Density Had a Minimal Effect on Overall Transduction Efficiency
Next, we investigated the influence of cell density on the transduction efficiency (Fig. 4). The concentrations of 40 and 60 PPC were chosen for this analysis as these concentrations had minimal impact on proliferation and prominent GFP expression levels. Overall, 60 PPC resulted in the highest expression levels for each cell density. For both virus concentrations, the 10,000 cells condition showed the slowest expression profile. The groups with 20,000 and 30,000 cells had a similar expression profile in the first phase of transduction, however, the maximum expression was achieved within 48 h for both virus concentrations using 20,000 cells/well (82.4% with 40 PPC and 89.7% with 60 PPC). Comparing the two viral loads at this density, 40 PPC had the lowest impact on confluency (data not shown). Representative time-lapse images of this condition are shown in Figure 4.
Discussion
The BacMam gene delivery system offers a number of different fluorescent fusion protein options to study mammalian cellular organelles and processes. The fluorescence expression can be analyzed using a fluorescent imager and conditions must be optimized to ensure high transduction efficiency with minimal cytotoxicity. This study demonstrated the ability of the CytoSMART Omni FL to optimize conditions for the transient transduction of HeLa cells with BacMam Nucleus-GFP constructs. The CytoSMART Omni FL allowed for high-throughput image acquisition and analysis of multiple transduction conditions with the ability to run long-term experiments.
We evaluated two different factors affecting transduction efficiency, namely cell concentration and virus concentration (particles per cell; PPC). Kinetic experiments showed that the transduction efficiency increased in a concentration-dependent manner to a maximum efficiency at 100 PPC.
We could also get a first impression on the cytotoxicity of the transduction using confluency data generated in the CytoSMART Cloud. All virus concentrations affected cell proliferation compared to the control group, with the highest confluence levels in 40 PPC groups and the lowest confluency with 100 PPC. The impact on cellular growth suggests that high viral loads, resulting in over-expression of Nucleus-GFP fusion protein, are toxic to HeLa cells [10].The second variable investigated in this study was the cell concentration. We found slight differences in GFP expression with the use of different cell densities. Maximum expression was achieved with 20,000 cells/well. Peak fluorescence levels were reached within 48 h; demonstrating the benefit of livecell imaging to determine the ideal incubation period. With the conducted measurements for GFP expression and least impact on confluency, we found that infection with 40 PPC at 20,000 cells was most suitable for the HeLa cell line.
Conclusion
In this study, we showed the application of high-throughput fluorescence live-cell imaging in optimizing transduction conditions of the BacMam system. The expression of Nucleus-GFP fusion proteins in HeLa cells could be kinetically monitored with the CytoSMART Omni FL and analyzed to calculate the percentage of GFP expressing cells as a function of virus concentration and cell density. Next to this, the integrated image analysis enabled automated brightfield confluence measurements as a non-invasive toxicity test. Consequently, we could identify the most optimal condition (40 PPC at a density of 20,000 cells/well) which resulted in high GFP expression with minimal effects on cell proliferation.
References
[1.] Kim, T. K. & Eberwine, J. H. (2010). Mammalian cell transfection: the present and the future. Anal. Bioanal. Chem. 397, 3173–3178.
[2.] Chong, Z. X., Yeap, S. K. & Ho, W. Y. (2021). Transfection types, methods and strategies: a technical review. PeerJ 9, e11165– e11165.
[3.] Hassan, N. J., Pountney, D. J., Ellis, C., & Mossakowska, D. E. (2006). BacMam recombinant baculovirus in transporter expression: a study of BCRP and OATP1B1. Protein expression and purification, 47(2), 591–598.
[4.] Grassi, G., Köhn, H., Dapas, B., Farra, R., Platz, J., Engel, S., Cjsareck, S., Kandolf, R., Teutsch, C., Klima, R., Triolo, G., & Kuhn, A. (2006). Comparison between recombinant baculo- and adenoviral-vectors as transfer system in cardiovascular cells. Archives of virology, 151(2), 255–271.
[5.] Ho, Y. C., Chung, Y. C., Hwang, S. M., Wang, K. C., & Hu, Y. C. (2005). Transgene expression and differentiation of baculovirustransduced human mesenchymal stem cells. The journal of gene medicine, 7(7), 860–868.
[6.] Hindriksen, S., Bramer, A. J., Truong, M. A., Vromans, M. J., Post, J. B., Verlaan-Klink, I., … & Hadders, M. A. (2017). Baculoviral delivery of CRISPR/Cas9 facilitates efficient genome editing in human cells. PloS one, 12(6), e0179514.
[7.] Abe, T., & Matsuura, Y. (2010). Host innate immune responses induced by baculovirus in mammals. Current gene therapy, 10(3), 226-231.
[8.] Mansouri, M., Bellon-Echeverria, I., Rizk, A., Ehsaei, Z., Cosentino, C. C., Silva, C. S., … & Berger, P. (2016). Highly efficient baculovirus-mediated multigene delivery in primary cells. Nature communications, 7(1), 1-13.
[9.] ThermoFisher Scientific (2021, December). BacMam Technology Overview. https://www.thermofisher.com/nl/en/home/ industrial/pharma-biopharma/drug-discovery-development/target-and-lead-identification-and-validation/pathway-biology/ cellular-pathway-analysis/bacmam-system/bacmam-technology-overview.html
[10.] Bolognesi B., Lehner B. (2018) Protein Overexpression: Reaching the limit. eLife 7:e39804.
Authors
Kim van Dongen, Application Scientist,
Inge Thijssen-van Loosdregt, Senior Application Scientist,
Mariya Yevstigneyeva, Scientific Content Manager
Application Science Department,
CytoSMART Technologies
Eindhoven, Netherlands